Gnaiger 2021 MitoFit BCA
| Gnaiger E (2021) Bioenergetic cluster analysis – mitochondrial respiratory control in human fibroblasts. MitoFit Preprints 2021.08. https://doi.org/10.26124/mitofit:2021-0008 |
» MitoFit Preprints 2021.08.
Bioenergetic cluster analysis – mitochondrial respiratory control in human fibroblasts
Gnaiger Erich (2021) MitoFit Prep
Abstract:
- Version 1 (v1) 2021-09-21 doi:10.26124/mitofit:2021-0008
Cell respiration reflects mitochondrial fitness and plays a pivotal role in health and disease. Despite the rapidly increasing number of applications of cell respirometry to address current challenges in biomedical research, cross-references are rare between respirometric projects and platforms. Evaluation of accuracy and reproducibility between laboratories requires presentation of results in a common format independent of the applied method. When cell respiration is expressed as oxygen consumption rate in an experimental chamber, normalization is mandatory for comparability of results. Concept-driven normalization and regression analysis are key towards bioenergetic cluster analysis presented as a graphical tool to identify discrete data populations.
In a meta-analysis of human skin fibroblasts, high-resolution respirometry and polarography covering cell senescence and the human age range are compared with multiwell respirometry. The common coupling control protocol measures ROUTINE respiration of living cells followed by sequential titrations of oligomycin, uncoupler, and inhibitors of electron transfer.
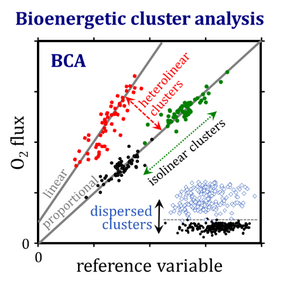
Bioenergetic cluster analysis increases the resolution of outliers within and differences between groups. An outlier-skewness index is introduced as a guide towards logarithmic transformation for statistical analysis. Isolinear clusters are separated by variations in the extent of a quantity that correlates with the rate, whereas heterolinear clusters fall on different regression lines. Dispersed clusters are clouds of data separated by a critical threshold value. Bioenergetic cluster analysis provides new insights into mitochondrial respiratory control and a guideline for establishing a quality control paradigm for bioenergetics and databases in mitochondrial physiology.
• Keywords: human dermal fibroblasts HDF, living cells ce, cell respiration, coupling control, oxidative phosphorylation OXPHOS, age, senescence, bioenergetic cluster analysis BCA, meta-analysis, normalization, high-resolution respirometry HRR, Oroboros O2k, Seahorse XF Analyzer, outlier-skewness index OSI, regression analysis • Bioblast editor: Gnaiger E • O2k-Network Lab: AT Innsbruck Oroboros
Data availability
- Original files are available Open Access at Zenodo repository: DOI 10.5281/zenodo.5518506
References
| Link | Reference | Year | View |
|---|---|---|---|
| Amrhein 2019 Nature | Amrhein V, Greenland S, McShane B (2019) Scientists rise up against statistical significance. Nature 567:305-7. | 2019 | PMID: 30894741 Open Access |
| Annesley 2019 Cells | Annesley SJ, Fisher PR (2019) Mitochondria in health and disease. Cells 8:680. doi: 10.3390/cells8070680 | 2019 | PMID: 31284394 Open Access |
| Curran-Everett 2020 Adv Physiol Educ | Curran-Everett D (2020) Evolution in statistics: P values, statistical significance, kayaks, and walking trees. Adv Physiol Educ 44:221-4. | 2020 | PMID: 32412384 Open Access |
| Dejmek 2018 Physiol Res | Dejmek J, Kohoutová M, Kripnerová M, Čedíková M, Tůma Z, Babuška V, Bolek L, Kuncová J (2018) Repeated exposure to hyperbaric hyperoxia affects mitochondrial functions of the lung fibroblasts. Physiol Res 67(Suppl 4):S633-S643. | 2018 | PMID: 30607970 Open Access |
| Doane 2011 J Statistics Education | Doane DP, Seward LE (2011) Measuring skewness: a forgotten statistic? J Statistics Education 19,2:1-18. | 2011 | Open Access |
| Doerrier 2018 Methods Mol Biol | Doerrier C, Garcia-Souza LF, Krumschnabel G, Wohlfarter Y, Mészáros AT, Gnaiger E (2018) High-Resolution FluoRespirometry and OXPHOS protocols for human cells, permeabilized fibers from small biopsies of muscle, and isolated mitochondria. Methods Mol Biol 1782:31-70. https://doi.org/10.1007/978-1-4939-7831-1_3 | 2018 | PMID: 29850993 » |
| Eisenhauer 2003 Teaching Statistics | Eisenhauer JG (2003) Regression through the origin. Teaching Statistics 25:76-80. | 2003 | https://onlinelibrary.wiley.com/doi/abs/10.1111/1467-9639.00136 |
| Gellerich 2004 Mitochondrion | Gellerich FN, Mayr JA, Reuter S, Sperl W, Zierz S (2004) The problem of interlab variation in methods for mitochondrial disease diagnosis: enzymatic measurement of respiratory chain complexes. Mitochondrion 4:427-39. | 2004 | PMID: 16120404 |
| Gnaiger 2001 Respir Physiol | Gnaiger E (2001) Bioenergetics at low oxygen: dependence of respiration and phosphorylation on oxygen and adenosine diphosphate supply. https://doi.org/10.1016/S0034-5687(01)00307-3 | 2001 | Respir Physiol 128:277-97. PMID: 11718759 |
| Gnaiger 2008 POS | Gnaiger E (2008) Polarographic oxygen sensors, the oxygraph and high-resolution respirometry to assess mitochondrial function. In: Mitochondrial dysfunction in drug-induced toxicity (Dykens JA, Will Y, eds) John Wiley & Sons, Inc, Hoboken, NJ:327-52. | 2008 | |
| Gnaiger 2020 BEC MitoPathways | Gnaiger E (2020) Mitochondrial pathways and respiratory control. An introduction to OXPHOS analysis. 5th ed. Bioenerg Commun 2020.2. https://doi.org/10.26124/bec:2020-0002 | 2020 | |
| Gnaiger 2020 MitoFit x | Gnaiger E (2021) The elementary unit — canonical reviewer's comments on: Bureau International des Poids et Mesures (2019) The International System of Units (SI) 9th ed. https://doi.org/10.26124/mitofit:200004.v2 | 2021 | MitoFit Preprints 2020.04.v2. The elementary unit — canonical reviewer's comments on: Bureau International des Poids et Mesures (2019) The International System of Units (SI) 9th ed. |
| BEC 2020.1 doi10.26124bec2020-0001.v1 | Gnaiger E et al ― MitoEAGLE Task Group (2020) Mitochondrial physiology. Bioenerg Commun 2020.1. https://doi.org/10.26124/bec:2020-0001.v1 | 2020 | Bioenerg Commun 2020.1. |
| Greco 2003 FASEB J | Greco M, Villani G, Mazzucchelli F, Bresolin N, Papa S, Attardi G (2003) Marked aging-related decline in efficiency of oxidative phosphorylation in human skin fibroblasts. FASEB J 17:1706-8. | 2003 | PMID: 12958183 Open Access |
| Han 2019 Front Physiol | Han Y, Zhou S, Coetzee S, Chen A (2019) SIRT4 and its roles in energy and redox metabolism in health, disease and during exercise. Front Physiol 10:1006. doi: 10.3389/fphys.2019.01006 | 2019 | PMID: 31447696 Open Access |
| Herst 2017 Front Endocrinol (Lausanne) | Herst PM, Rowe MR, Carson GM, Berridge MV (2017) Functional mitochondria in health and disease. Front Endocrinol (Lausanne) 8:296. doi: 10.3389/fendo.2017.00296. | 2017 | PMID: 29163365 Open Access |
| Hood 2019 Annu Rev Physiol | Hood DA, Memme JM, Oliveira AN, Triolo M (2019) Maintenance of skeletal muscle mitochondria in health, exercise, and aging. Annu Rev Physiol 81:19-41. doi: 10.1146/annurev-physiol-020518-114310 | 2019 | PMID: 30216742 |
| Huetter 2004 Biochem J | Hütter E, Renner K, Pfister G, Stöckl P, Jansen-Dürr P, Gnaiger E (2004) Senescence-associated changes in respiration and oxidative phosphorylation in primary human fibroblasts. https://doi.org/10.1042/BJ20040095 | 2004 | Biochem J 380:919-28. PMID: 15018610 - Open Access |
| Jaber 2020 BM Exp Neurol | Jaber SM, Yadava N, Polster BM (2020) Mapping mitochondrial respiratory chain deficiencies by respirometry: beyond the Mito Stress Test. BM Exp Neurol 328:113282. | 2020 | PMC7202675 Open Access |
| Kahneman 2011 Penguin Books | Kahneman D (2011) Thinking, fast and slow. Penguin Books 499 pp. | 2011 | Penguin |
| Karabatsiakis 2014 Transl Psychiatry | Karabatsiakis A, Boeck C, Salinas-Manrique J, Kolassa S, Calzia E, Dietrich DE, Kolassa IT (2014) Mitochondrial respiration in peripheral blood mononuclear cells correlates with depressive subsymptoms and severity of major depression. Transl Psychiatry 4:e397. | 2014 | PMID: 26126180 Open Access » |
| Kozieł 2011 J Invest Dermatol | Kozieł R, Greussing R, Maier AB, Declercq L, Jansen-Dürr P (2011) Functional interplay between mitochondrial and proteasome activity in skin aging. J Invest Dermatol 131:594-603. | 2011 | PMID: 21191400 Open Access |
| Kuffner 2020 Cells | Kuffner K, Triebelhorn J, Meindl K, Benner C, Manook A, Sudria-Lopez D, Siebert R, Nothdurfter C, Baghai TC, Drexler K, Berneburg M, Rupprecht R, Milenkovic VM, Wetzel CH (2020) Major depressive disorder is associated with impaired mitochondrial function in skin fibroblasts. Cells 9:884. doi: 10.3390/cells9040884 | 2020 | PMID: 32260327 Open Access |
| Mahapatra 2018 Clin Sci (Lond) | Mahapatra G, Smith SC, Hughes TM, Wagner B, Maldjian JA, Freedman BI, Molina AJA (2018) Blood-based bioenergetic profiling is related to differences in brain morphology in African Americans with Type 2 diabetes. Clin Sci (Lond) 132:2509-18. https://doi.org/10.1042/CS20180690 | 2018 | PMID: 30401689 Open Access |
| Memme 2021 J Physiol | Memme JM, Erlich AT, Phukan G, Hood DA (2021) Exercise and mitochondrial health. J Physiol 599:803-17. doi: 10.1113/JP278853 | 2021 | PMID: 31674658 |
| Miettinen 2017 Trends Cell Biol | Miettinen TP, Björklund M (2017) Mitochondrial function and cell size: an allometric relationship. Trends Cell Biol 27:393-402. | 2017 | PMID: 28284466 |
| Nunnari 2012 Cell | Nunnari J, Suomalainen A (2012) Mitochondria: in sickness and in health. Cell 148:1145-59. https://doi.org/10.1016/j.cell.2012.02.035 | 2012 | PMID: 22424226 Open Access |
| Renner 2003 Biochim Biophys Acta | Renner K, Amberger A, Konwalinka G, Gnaiger E (2003) Changes of mitochondrial respiration, mitochondrial content and cell size after induction of apoptosis in leukemia cells. Biochim Biophys Acta 1642:115-23. | 2003 | PMID: 12972300 Open Access |
| Savage 2007 Proc Natl Acad Sci U S A | Savage VM, Allen AP, Brown JH, Gillooly JF, Herman AB, Woodruff WH, West GB (2007) Scaling of number, size, and metabolic rate of cells with body size in mammals. Proc Natl Acad Sci U S A 104:4718-23. | 2007 | PMID: 17360590 Open Access |
| Schoepf 2020 Nat Commun | Schöpf Bernd, Weissensteiner Hansi, Schäfer Georg, Fazzini Federica, Charoentong Pornpimol, Naschberger Andreas, Rupp Bernhard, Fendt Liane, Bukur Valesca, Giese Irina, Sorn Patrick, Sant’Anna-Silva Ana Carolina, Iglesias-Gonzalez Javier, Sahin Ugur, Kronenberg Florian, Gnaiger Erich, Klocker Helmut (2020) OXPHOS remodeling in high-grade prostate cancer involves mtDNA mutations and increased succinate oxidation. https://doi.org/10.1038/s41467-020-15237-5 | 2020 | Nat Commun 11:1487. PMID: 32198407 Open Access » |
| Silver 2012 Penguin Press | Silver N (2012) The signal and the noise. The art and science of prediction. Penguin Press:534 pp. | 2012 | |
| Steinlechner-Maran 1996 Am J Physiol Cell Physiol | Steinlechner-Maran R, Eberl T, Kunc M, Margreiter R, Gnaiger E (1996) Oxygen dependence of respiration in coupled and uncoupled endothelial cells. Am J Physiol Cell Physiol 271:C2053-61. https://doi.org/10.1152/ajpcell.1996.271.6.C2053 | 1996 | PMID: 8997208 |
| Ulrich 2020 Elife | Ulrich R, Miller J (2020) Meta-research: Questionable research practices may have little effect on replicability. Elife 9:e58237. doi: 10.7554/eLife.58237 | 2020 | PMID: 32930092 Open Access |
| Villani 1997 Proc Natl Acad Sci U S A | Villani G, Attardi G (1997) In vivo control of respiration by cytochrome c oxidase in wild-type and mitochondrial DNA mutation-carrying human cells. Proc Natl Acad Sci U S A 94:1166-71. | 1997 | PMID: 9037024 Open Access |
| Wagner 2011 Free Radic Biol Med | Wagner BA, Venkataraman S, Buettner GR (2011) The rate of oxygen utilization by cells. Free Radic Biol Med 51:700-12. | 2011 | |
| Wallace 2010 Annu Rev Pathol | Wallace DC, Fan W, Procaccio V (2010) Mitochondrial energetics and therapeutics. Annu Rev Pathol 5:297-348. | 2010 | PMID: 20078222 |
| Wilcox R 2015 Journal of Statistical Distributions and Applications | Wilcox RR (2015) Comparing the variances of two dependent variables. Journal of Statistical Distributions and Applications 2:7. doi:10.1186/s40488-015-0030-z | 2015 | Journal of Statistical Distributions and Applications ResearchGate |
| Wu 2007 Am J Physiol Cell Physiol | Wu M, Neilson A, Swift AL, Moran R, Tamagnine J, Parslow D, Armistead S, Lemire K, Orrell J, Teich J, Chomicz S, Ferrick DA (2007) Multiparameter metabolic analysis reveals a close link between attenuated mitochondrial bioenergetic function and enhanced glycolysis dependency in human tumor cells. https://doi.org/10.1152/ajpcell.00247.2006 | 2007 | Am J Physiol Cell Physiol 292:C125-36. PMID: 16971499 Open Access |
| Yepez 2018 PLOS One | Yépez VA, Kremer LS, Iuso A, Gusic M, Kopajtich R, Koňaříková E, Nadel A, Wachutka L, Prokisch H, Gagneur J (2018) OCR-Stats: Robust estimation and statistical testing of mitochondrial respiration activities using Seahorse XF Analyzer. https://doi.org/10.1371/journal.pone.0199938 | 2018 | PLOS ONE 13:e0199938. PMID: 29995917 Open Access |
| Zdrazilova 2022 PLOS ONE | Zdrazilova L, Hansikova H, Gnaiger E (2022) Comparable respiratory activity in attached and suspended human fibroblasts. PLoS ONE 17:e0264496. https://doi.org/10.1371/journal.pone.0264496 | 2022 | PMID: 35239701 Open Access |
| Zhang 2021 PLOS ONE | Zhang X, Yuan T, Keijer J, de Boer VCJ (2021) OCRbayes: a Bayesian hierarchical modeling framework for Seahorse extracellular flux oxygen consumption rate data analysis. PLOS ONE 16:e0253926. | 2021 | PMID: 34265000 Open Access |
| De Mello 2018 Life Sci | de Mello AH, Costa AB, Engel JDG, Rezin GT (2018) Mitochondrial dysfunction in obesity. Life Sci 192:26-32. doi: 10.1016/j.lfs.2017.11.019 | 2018 | PMID: 29155300 |
Living communication: additional references
- Aisenberg AC (1961) The glycolysis and respiration of tumors. Academic Press 224 pp. - »Bioblast link« - Extensive compilation of respirometric data on tissue slices and cells expressed as volume of O2 per hour per dry mass, based mainly on manometric respirometry, including anaerobic and aerobic glycolysis (Pasteur effect), O2 consumption and CO2 production.
- Stahel WA (2021) New relevance and significance measures to replace p-values. PLOS ONE 16:e0252991. https://doi.org/10.1371/journal.pone.0252991.
Cited by
- Zdrazilova L, Hansikova H, Gnaiger E (2021) Comparable respiratory activity in attached and suspended human fibroblasts. MitoFit Preprints 2021.7. doi:10.26124/mitofit:2021-0007 - »Bioblast link«
- Krako Jakovljevic N, Ebanks B, Katyal G, Chakrabarti L, Markovic I, Moisoi N (2021) Mitochondrial homeostasis in cellular models of Parkinson’s Disease. Bioenerg Commun 2021.2. https://doi.org/10.26124/bec:2021-0002
- Komlódi T, Cardoso LHD, Doerrier C, Moore AL, Rich PR, Gnaiger E (2021) Coupling and pathway control of coenzyme Q redox state and respiration in isolated mitochondria. Bioenerg Commun 2021.3. https://doi.org/10.26124/bec:2021-0003
Acknowledgements version 1
- I thank Lucie Zdrazilova for collaboration and input from our parallel manuscript, and Hans Zischka, Pablo M Garcia-Roves, Omar Torres-Quesada, Anthony JA Molina, Philip A Kramer, Jenny L Gonzalez-Armenta, Mateus Grings, and the Oroboros team ― particularly Chris Donnelly, Sabine Schmitt, Timea Komlódi, Carolina Doerrier, and Luiza HD Cardoso ― for stimulating discussions and critical comments.
- To promote transparency, MitoFit Preprints encourages Open Access to relevant scientific contributions of acknowledged persons, even though it may not be possible to provide a fully balanced account of the correspondence and personal discussions.
- Lucie Zdrazilova: see Zdrazilova L, Hansikova H, Gnaiger E (2021) Comparable respiratory activity in attached and suspended human fibroblasts. MitoFit Preprints 2021.7. doi:10.26124/mitofit:2021-0007
- Chris Donnelly (v0.1 2021-06-15): One important point I believe is worthy to mention here is that in the Yépez paper they did not titrate Ama. I believe Rox is after Rot titration.
- Sabine Schmitt (v0.1 2021-06-15): The term concept-driven normalization requires explanation.
- Timea Komlódi (v0.1 2021-06-15): I found it very useful how you explain the different outlier levels and use these for excluding data. Regarding negative values, sometimes we observe negative O2 flux in the ROX state which is usually used for baseline correction. Can we use this “negative Rox” for correction which would lead to higher O2 fluxes? Or should we exclude those files where Rox is negative? Or in this case instead of baseline-corrected O2 flux we can only use flux control ratios in the whole study?
- Hans Zischka (v0.2 2021-06-16): I would separate the discussion and conclusions. The conclusions are at best unclear.
- Pablo M Garcia-Roves (v0.2 2021-06-16): I like the topic of the manuscript but I think it needs additional work to get the message deliver clearly. In my opinion it needs a different structure where you clearly define sequential steps in your rational and analysis. For example, Figure 1 address several issues at the same time and it could be confusing at the time to assess data representation. One part of the manuscript is dedicated to explain how data analysis has been performed (very important and informative: PSI, OSI, log-transformed; BCA, …). But explanations about the procedure to perform data analysis are intercalated with data representation and analysis. The work brings to my attention a way to analyze data sets that could be of interest, as you mentioned in the manuscript, for data collected during the MitoEAGLE COST action.
- Anthony JA Molina (version 0.4 2021-06-24): Take a look at this paper that just came out (attached) about the same dataset. - Abstract: Maybe "quality control paradigm for bioenergetics..." - I think you should also add a paragraph about limitations of BCA and what types of this studies this would be relevant to. For example, exclusion of data can be an issue in certain types of clinical studies. - I would add that the selection of the reference for normalization is driven by the question being asked by a particular study.
- Philip A Kramer (version 0.4 2021-06-24): Worth noting that (Mahapatra) Gargi's paper was not a comparison of data from two different laboratories. {Jenny: Also, in Gargi's paper the Seahorse data and the O2k data are treated as separate but complementary entities and aren't compared to each other.} - Might be worth noting here that cell volume can be broken down even further to cytoplasmic volume where the mitochondria reside. This is not captured by total protein. This is especially important when dealing with mixed cell populations like PBMCs as lymphocytes have a nuclear to cytoplasmic ratio of about 4:1 vs monocytes of about 2:1. Contaminating platelets in these isolations have no nucleus at all. N:C ratios can vary widely based on maturity and degree of activation. - Edge effects are the only explanation provided for this and we don't know whether this is an isolated artifact of a single machine, user, or of the seahorse platform itself, or contamination or leaking with antimycin, oligo, etc during mixing steps, or microbubbles against the sensor. Might be worth mentioning limitations of the BCA in this single data set. I would like to know how many seahorse plates, i.e. independent runs all of these data points cover. Is each n a well? {Erich: Yes, this is correct.} - If 2822 data points, are we talking about 30 different seahorse plates? {Erich: There were 124 different plates with a median of 20 wells per plate. The effect of averaging wells per plate is shown in new Figure 8 (v0.12).} - Did you exclude all edges on each plate to test for the edge effects? {Erich: Yes, all edges were excluded in the test.} - Seems to me that BCA is only applicable to Seahorse datasets as it is unlikely anyone will ever have this many data points for O2k. For this to be a comparison paper, consider discussing both instruments in each section, even if just to say BCA isn't applicable for O2k. {Erich: BCA is applicable to small and large data sets, as shown in several figures, such as Figure 7 for small data sets and Figure 8 comparing a large and small data set directly.} - It might be worth comparing O2k data with 3 long read intervals that correspond to the measurement cycles of a Seahorse and seeing if this is a platform issue or a timing issue. {Erich: The standard procedures were used for each platform for comparison.}
- Jenny L Gonzalez-Armenta (version 0.4 2021-06-24): Intro: ..in many other tissues and in mito dysfunction associated with many other conditions - aging, diabetes, etc. Are these the best data sets to use? Here are my concerns: (1) I have issues with the quality of the Yepez data. I truly believe the Seahorse data is not the best representation of the kind of data you would want to use. (2) While the plate set up is quite nice and there are plenty of technical replicates, there is so much variability even in their NHDF control cell line. The basal respiration for those replicates across plates range from ~6 - 186 pmol/min which is very large and not at all what I would expect from carefully controlled experiments. In my hands, I have very little plate to plate variability with controls - though they are a different cell type. (3) They make no mention of any steps that they took to minimize plate to plate variability - in fact, there are no details about their culture methods. Therefore, I have to assume they did not do anything special in terms of culture - was it carefully controlled for batch to batch effects of media, supplements (we have seen a large bioenergetic difference based on FBS batches), passage number, how they were stored, etc. Likely, these have a greater effect along with the user than the instrumentation. (4) I would expect a lot of variability from fibroblasts derived from mito disease patients - I hope they were not included in the analysis. (5) I am much more comfortable with the quality of the data from the Hutter paper. The methods are clearly laid out and passage numbers are included. However, I am concerned about the statistical implications of comparing 12 replicates from O2k to thousands of replicates from the Seahorse data. (6) Overall, I would be much more comfortable with an instrumental comparison that controlled for differences in cell line, culture methods, passage number, facility, lab personnel, etc. so that we can be sure that it is a true comparison and not a measurement of these differences. {Erich: See the parallel paper, Zdrazilova et al 2021.} - Is the difference in recording interval something that should be considered in the analysis? Does this limit the ability to compare the platforms? Just curious about that from a statistical standpoint. - Again, this makes me question their culture methods. I really don't believe that the cells are consistently healthy between plates and the very low OCR suggest to me that they did not passage after thaw or are using high passage numbers.
- Carolina Doerrier (version 0.4 2021-06-24): The most difficult part to follow was the OSI and the most stimulating sections those showing the bioenergetic clusters. At the beginning of section 3.2, I miss a brief explanation of why PSI was modified in the OSI index and, what is the OSI index. I find it easier to understand following the information from the website (Outlier-skewness index), maybe you could add a similar brief explanation in the manuscript and why this index is better than PSI. From where is coming the critical value indicative of outliers |OSI|>0.03? In Fig. 1, I would suggest the same scale for Fig 1b and Fig 1d. In Figure 5a you represented R3 as a function of E (shown in the graph). I would suggest to add it (R3) also in the figure legend. Maybe you could add “(bc)” in the label of the axis when it is baseline corrected for Rox. Why the different measurements in ROUTINE, LEAK, and ET are not in italic (e.g. R1, R2 …)? Is there a reason? The cluster are in italic (e.g. LC2), but not the different measurements (e.g. L1, L2, L3). In figure 6, are the measurements in LC3 the same than the measurements in ROXC2? By removing the rC2 cluster (line 476, and figure 7d, e and f), the cluster LC3 is out. Do these measurements correspond to ROXC2 as well? In figure 7, is the cluster lC2 is written like that just to distinguish it from LC2? Once you read all figure it is very easy to understand and follow it. The values from the cluster lC2 (with low coupling efficiency threshold (E-L)/E<0.8) are eliminated (Fig. 7b). In ROUTINE R3, the values which correspond to the same measurements (lC2) are not eliminated (Fig. 7b), however, we see that they differ from rC1 (and the same measurements have lower coupling efficiency). Would not be correct to eliminate these measurements as well? How do you define the threshold values for outlier exclusion (Figure 7)? If people will apply the BCA for analysis of the own data, I think it will be useful to provide a short explanation. In all axis of the graphs (including FCR or coupling control efficiencies) maybe could be good to add which measurement is used, for example in Figure 7e, (R3-Lmin/E1) net control ratio. In figure 8, maybe you could add in the graphs the instrument used, something similar like you have in Figure 1. Slopes and intercept are not clear for me in figure 8.
- Omar Torres-Quesada (version 0.4 2021-06-24): Concerning the threshold values for detection of outliers (Figure 7, page 13), are these values specifically calculated for each data set or are they standard? This would be quite interesting to know if someone wants to use this analysis which own data. The BCA outlier level workflow shows a stepwise strategy to remove outliers from large-scale data. Would it be possible to apply this strategy with small-scale data? This would be relevant for example for BCA of O2k data (usually with lower population size). The excel file I find it quite useful when somebody wants to apply the same workflow with own data.
- Mateus Grings (v0.9, 2021-08-30): I really liked the ideas and data of the manuscript and agree that bioenergetic cluster analysis is a very interesting approach to analyze and compare bioenergetics data from different cells and performed with different instruments. Besides that, it was importantly pointed out that it is a great tool that can help in the evaluation of reproducibility of data. It is interesting that since it is a more visual approach, it is easier to observe differences among different data and also for the efficient detection of outliers. It was very easy and transparent to visualize the outliers and understand the problems related to them in the graphical analyzes performed for their characterization (Figures 9 and 10). I liked the general idea of normalization using internal experimental values for the comparison of data, excluding errors that may be induced by the addition of external values, such as differences in procedures to measure the external parameters in distinct laboratories. In addition, the use of normalization with flux control ratios and flux control efficiencies may show different aspects of the data. Although I do not have experience with cluster analysis and profound knowledge of all the mathematical concepts used to develop this methodology, the conceptual background was very clear and helpful for the understanding of the manuscript results. I did not profoundly understand all details in some of the results, but I could understand the main results and ideas of all of them, especially because of my previous knowledge on respirometry. I also found particularly interesting the models used in Figure 12 to observe the differentiation between the dyscapacity of fibroblasts from aged versus young donors and dyscoupling in senescent fibroblasts versus young proliferating and growth-arrested. Some problems pointed out about experiments performed with the Seahorse XF96 instrument were very relevant. Something that was mentioned and that I always think about is the use of a single uncoupler injection and the lack of uncoupler titration. Even though ideal concentrations of uncoupler for measuring ET capacity are tested in a pilot experiment for different cell types, the conditions of the cells or the assay may change in different experiments and situations, so that the uncoupler concentration needed for efficiently measuring ET capacity may vary (making it important to do a titration for each experiment). Another matter that I would like to comment about regards the normalization by cell number at the Seahorse XF96. I thought about it when I was reading the methodology used for NHDF in the Seahorse (normalization by seed count versus final cell count). When I performed some of my experiments in the Seahorse using fibroblasts from different patients with the same disease, I could not normalize the data using the seed count. This happened because the cells had very different proliferation rates and showed different adhesion patterns after seeding due to differences in patient genotypes and phenotypes. Therefore, I had to evaluate the cells after the assay to do the normalization. I think it is important to take this into account when performing normalization of experiments with attached cells.- Figure 3: Wasn’t Rox measured in the presence of Rot&Ama, instead of Rot or Rot&Ama? {Erich: In Hütter et al (2004), Rox was measured sequentially by titration or Rot (evaluated as a sparate stead state) followed by Ama. No difference was observed after titration of Ama, and Rox was defined as the minimum value in either state ROX1 (Rot) or ROX2 (Rot&Ama). In Yépez et al (2018), most ROX states are specified as Rot&Ama, but some are shown as Rot. The data from Hütter et al (2004) suggest, that the difference can be ignored.} - According to BCA outlier level 2, 60 outliers belonging to cluster lC2 were excluded, and according to BCA outlier level 3, 113 outliers belonging to cluster rC2 were excluded. This gives a total of 173 eliminated data. However, it is mentioned that 2 data points belong to both cluster lC2 and rC2. So, my question is: were these 2 common data points initially excluded within the 60 outlier of cluster lC2 and are not included in the 113 outliers excluded in the cluster rC2, so that the total of excluded data is 173? Or do the 60 and 113 outliers of the 2 different clusters contain these 2 common data points? {Erich: Thanks for pointing this out. A final check revealed that there were 9 physiological outliers, 62 wells were in cluster lC2 and 115 wells were in cluster rC2 with two wells overlapping (thus 175 outliers were eliminated for lC2 and rC2 together), and 8 statistical outliers remained. Taken together, 192 outliers were eliminated from the data set with 2822 wells, such that 2630 wells were used for the final analysis.}
- I thank Luiza HD Cardoso for helpful discussion on the data repository zenodo.
Acknowledgements towards version 2
- I thank Bengt Kayser for bringing to my attention the concept of perpendicular regression.
Support
- This work was partially funded by the European Union’s Horizon 2020 research and innovation programme under grant agreement No. 859770, NextGen-O2k project. Contribution to the MitoEAGLE Task Group of the Mitochondrial Physiology Society.
Keywords
 Keywords—MitoPedia - >>>>>>> - Click on [Expand] or [Collapse] - >>>>>>>
Keywords—MitoPedia - >>>>>>> - Click on [Expand] or [Collapse] - >>>>>>>
Labels: MiParea: Respiration, Instruments;methods
Pathology: Aging;senescence
Organism: Human Tissue;cell: Fibroblast Preparation: Intact cells
Regulation: Coupling efficiency;uncoupling Coupling state: LEAK, OXPHOS, ET Pathway: ROX HRR: Oxygraph-2k
Comparison of respirometric methods, MitoEAGLEPublication, SUIT-003, MitoFit 2021 CoQ, PLoSONE2022ace-sce, MitoFit2022QC
| Term | Link to MitoPedia term | Symbol | Unit | Links and comments |
|---|---|---|---|---|
| catabolic rate of respiration | Cell respiration | JkO2; IkO2 | varies | flux J versus flow I |
| catabolic reaction | Cell respiration | k | - | |
| cell count | Count | Nce | [x] | see number of cells; countable object s=ce |
| cell-count concentration | Concentration | Cce | [x∙L-1] | Cce = Nce∙V-1; count concentration C versus amount concentration c; subscript ce indicates the entity type: concentration of ce. But it does not signal 'per entity', which would be written as 'per cell' Xce. |
| cell mass | Body mass | mce | [kg] | mass of cells m versus mass per cell (per single entity cell) MXce |
| cell mass, mass per cell | Body mass | MXce | [kg∙x-1] | mass per single cell MXce; upper case M and subscript X signal 'per count', subscript ce signals the entity s=ce; in a context restricted to cells or molecules or a particular organism such as humans, the abbreviated symbol M [kg∙x-1] provides a sufficiently informative signal, particularly in combination with the explicit unit. |
| cell-mass concentration in chamber | Concentration | Cmce | [kg∙L-1] | see Cms: Cmce = mce∙V-1; upper case C alone would signal 'count concentration' (CN is more explicit), whereas the signal for 'mass concentration' is in the combination Cm. |
| concentration of O2, amount | Concentration | cO2 = nO2∙V-1 | [mol∙L-1] | [O2] |
| concentration of s, count | Concentration | Cs = Ns∙V-1 | [x∙L-1] (number concentration Cohen 2008 IUPAC Green Book); the signal for count concentration is given by the upper case C in contrast to c for amount concentration. In both cases, the subscript X indicates the entity type, not to be confused with a number of entities. | |
| count of Xs | Count | Ns | [x] | SI; see number of entities Xs |
| coupling control | Coupling-control ratio | CCR | - | |
| coupling control state | Coupling control state | CCS | - | |
| electron transfer pathway | Electron transfer pathway | ET pathway | - | |
| electron transfer, state | Electron transfer pathway | ET | - | (State 3u) |
| electron transfer system | Electron transfer pathway | ETS | - | (electron transport chain) |
| elementary entity | Entity | Xs | [x] | single countable object of sample type s |
| ET capacity | ET capacity | E | varies | rate |
| flow, for O2 | Flow | IO2 | [mol∙s-1] | system-related extensive quantity |
| flux, for O2 | Flux | JO2 | varies | size-specific quantity |
| flux control ratio | Flux control ratio | FCR | 1 | background/reference flux |
| International System of Units | International System of Units | SI | - | Cohen 2008 IUPAC Green Book |
| LEAK state | LEAK respiration | LEAK | - | (compare State 4) |
| LEAK respiration | LEAK respiration | L | varies | rate |
| living cells | Living cells | ce | - | (intact cells) |
| mass concentration of sample s in chamber | Concentration | Cms | [kg∙L-1] | |
| mass of sample s in a mixture | Mass | ms | [kg] | SI: mass of pure sample mS |
| mass per single object | Body mass | MNX | [kg∙x1] | SI: m(X); compare molar mass M(X) |
| mitochondria or mitochondrial | Mitochondria | mt | - | |
| mitochondrial concentration | Mitochondrial marker, Concentration | CmtE = mtE∙V-1 | [mtEU∙L-1] | |
| mitochondrial content per X | Mitochondrial marker | mtENX | [mtEU∙x-1] | mtENX = mtE∙NX-1 |
| mitochondrial elementary marker | Mitochondria | mtE | [mtEU] | quantity of mt-marker |
| mitochondrial elementary unit | Mitochondria | mtEU | varies | specific units for mt-marker |
| MitoPedia | MitoPedia, MitoPedia: Respiratory states | |||
| normalization of rate | Normalization of rate | - | - | |
| number of cells | Count | Nce | [x] | total cell count of living cells, Nce = Nvce + Ndce |
| oxidative phosphorylation | Oxidative phosphorylation | OXPHOS | - | |
| OXPHOS-capacity | OXPHOS-capacity | P | varies | rate |
| OXPHOS state | OXPHOS-capacity | OXPHOS | - | OXPHOS-state distinguished from the process OXPHOS (State 3 at kinetically-saturating [ADP] and [Pi]) |
| oxygen concentration | Oxygen concentration | cO2 = nO2∙V-1 | [mol∙L-1] | [O2] |
| oxygen solubility | Oxygen solubility | SO2 | [µmol·kPa-1] | |
| oxygen flux, in reaction r | Oxygen flux | JrO2 | varies | |
| quantities, symbols, and units | Quantities, symbols, and units | - | - | An explanation of symbols and unit [x] |
| rate in ET state | Electron transfer pathway | E | varies | ET capacity |
| rate in LEAK state | LEAK respiration | L | varies | L(Omy) |
| rate in ROX state | Residual oxygen consumption | Rox | varies | |
| residual oxygen consumption | Residual oxygen consumption | ROX; Rox | - | state ROX; rate Rox |
| respiration | Respirometry | JrO2 | varies | rate of reaction r |
| respiratory state | MitoPedia: Respiratory states | - | - | |
| steay state | Steady state | - | - | |
| substrate-uncoupler-inhibitor-titration | Substrate-uncoupler-inhibitor titration | SUIT | - | |
| system | System | - | - | |
| unit elementary entity | Entity | UX | [x] | single countable object |
| uncoupling | Uncoupler titrations | - | - | |
| volume of experimental chamber | Volume | V | [L] | liquid volume V including the sample s |