The printable version is no longer supported and may have rendering errors. Please update your browser bookmarks and please use the default browser print function instead.
Description
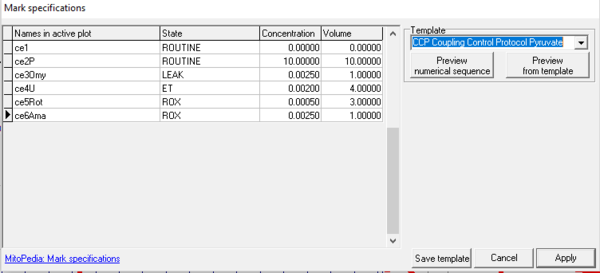
The function Mark specifications is largely replaced by SUIT DL-Protocols and Instrumental DL-Protocols in DatLab 7.4. Mark specifications allow the user to rename Marks in the active plot and save/recall the settings. Rename marks individually by clicking into the horizontal bar, or use corresponding templates for renaming the entire sequence of marks.
- Mark specifications (name, state, concentration and volume) are shown from the active plot. Define the template name and save template.
- Select a Template, left click on preview from template, and rename all marks of the plot, including the state, concentration and volume.
Reference mark names in DatLab 7
| Reference mark names | Description | SUIT protocol | Reference |
|---|---|---|---|
| MiPNet08.09_IntactCells | Coupling control protocol (ceCCP) in intact cells. | 1ce;2ceOmy;3ceU- | MiPNet08.09 CellRespiration
|
| MiPNet10.04_IntactCells | Coupling control protocol (ceCCP) in intact cells using TIP2k for automatic uncoupler titrations. | 1ce;2ceOmy;3ceU- | MiPNet10.04 CellRespiration
|
| MiPNet21.06 RP1pce | |||
| MiPNet21.06 RP2pce | |||
| MiPNet21.06 RP1mt | |||
| MiPNet21.06 RP2mt |
- » Library of SUIT protocols MitoPedia: SUIT protocols
Keywords
- Bioblast links: Marks in DatLab - >>>>>>> - Click on [Expand] or [Collapse] - >>>>>>>
- Specific
- O2k-Procedures
- MiPNet O2k-Procedures
- » MiPNet26.06 DatLab 7: Guide - Section on setting Marks
- MiPNet O2k-Procedures
- General
MitoPedia O2k and high-resolution respirometry:
DatLab