Jusic 2019 MitoFit Preprint Arch EA
| Jusic Amela, Hajrulahovic A, Devaux Y (2019) Noncoding RNAs regulatory network in mitochondria. https://doi.org/10.26124/mitofit:ea19.MiPSchool.0006 |
» MitoFit Preprint Arch EA19.6.
Noncoding RNAs regulatory network in mitochondria
Jusic A, Hajrulahovic A, Devaux Y (2019) MitoFit Preprint Arch
Abstract: Version 1 (v1) 2019-06-03 doi:10.26124/mitofit:ea19.MiPSchool.0006
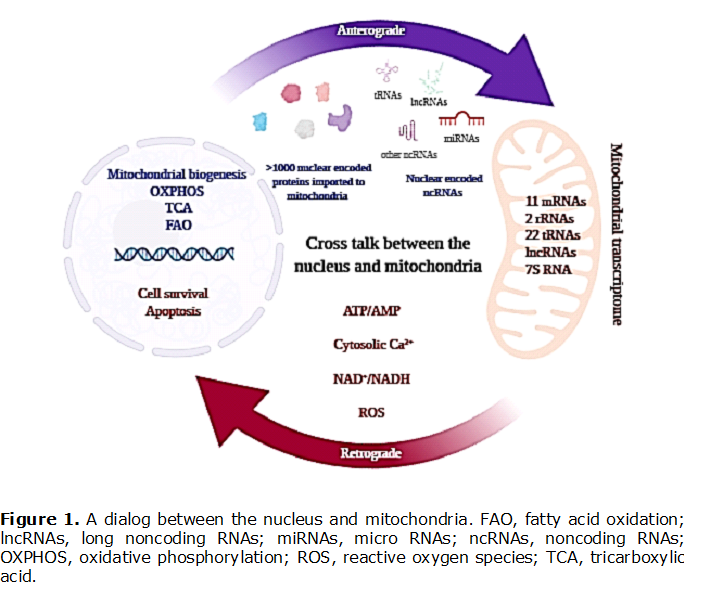
Mitochondria are powerhouses of eukaryotic cells and possess own bioenergetically specialized genetic system also known as mitochondrial DNA (mtDNA). In human cells, mtDNA encodes 22 transfer RNAs, 2 ribosomal RNAs and 13 proteins which are subunits of the oxidative phosphorylation machinery. Due to the small size of the mitochondrial genome, mitochondrial biogenesis and function are largely dependent on a number of nuclear-encoded molecules that are imported to mitochondria via the translocase of the outer mitochondrial membrane complex. In line with that, the mitochondrial proteome and transcriptome landscape represents an intricate mixture of intrinsic (i.e. mitochondrial) and extrinsic (i.e. nuclear-encoded) molecules. Thus, mitochondrial metabolism, as well as cellular homeostasis, require coordination of expression of two genomes and well-tuned cross-talk between the nucleus and mitochondria [Figure 1]. Although significant progress has been achieved in elucidating the molecular mechanisms of mito-nuclear communication, the regulatory network of this communication remains to be fully explored. Mitochondria are able to import various RNAs from the cytosol and also export particular RNAs. An increasing body of data from genome-wide analyses and RNA sequencing experiments has revealed that a large part of the human genome is transcribed into RNA molecules that are unable to encode proteins, the so-called noncoding RNAs (ncRNAs). Arbitrarily, ncRNAs are classified into small ncRNAs, which are up to 200 nucleotides long, and long ncRNAs RNAs (lncRNAs), which are longer than 200 nucleotides. Small RNAs contain, the well-known microRNAs (miRNAs) which down-regulate the expression of target messenger RNAs through base pairing. LncRNAs regulate gene expression mostly at the epigenetic level. According to the last GENCODE release (v30), the human genome contains 7576 small and 16193 long noncoding RNA genes. Accumulating evidence indicates that ncRNAs may contribute to the synchronization of essential cellular and mitochondrial biological pathways, acting as "messengers" between the mitochondria and the nucleus [1]. - Extended abstract • Keywords: mtDNA • Bioblast editor: Iglesias-Gonzalez J
Affiliations
Jusic Amela(1), Hajrulahovic A(1), Devaux Y(2)
- Dept of Biology, Faculty of Natural Sciences and Mathematics, Univ of Tuzla, Bosnia and Herzegovina - amela.jusic@untz.ba
- Cardiovascular Research Unit, Luxembourg Institute of Health, Luxembourg, Luxembourg
Results
References
- Vendramin R, Marine JC, Leucci E (2017) Non-coding RNAs: the dark side of nuclear-mitochondrial communication. EMBO J 36:1123-1133.
- Goretti E, Wagner DR, Devaux Y (2014) miRNAs as biomarkers of myocardial infarction: a step forward towards personalized medicine? Trends Mol Med. 20:716-725.
- Steer CJ (2015) microRNAs in Mitochondria: An Unexplored Niche. Adv Exp Med Biol 887:31-51.
- Duarte FV, Palmeira CM, Rolo AP (2014) The Role of microRNAs in Mitochondria: Small Players Acting Wide. Genes (Basel) 5:865–886.
- Murri M, El Azzouzi H (2018) MicroRNAs as regulators of mitochondrial dysfunction and obesity. Am J Physiol Heart Circ Physiol 1;315:H291-H302.
- Rackham O, Shearwood AM, Mercer TR, Davies SM, Mattick JS, Filipovska (2011) A Long noncoding RNAs are generated from the mitochondrial genome and regulated by nuclear-encoded protein. RNA 17:2085–2093.
- Leucci E, Vendramin R, Spinazzi M, Laurette P, Fiers M, Wouters J, Radaelli E, et al. (2016) Melanoma addiction to the long non-coding RNA SAMMSON. Nature 531:518-522.
- Kumarswamy R, Bauters C, Volkmann I, Maury F, Fetisch J, Holzmann A, et al. (2014) Circulating long noncoding RNA, LIPCAR, predicts survival in patients with heart failure. Circ Res 114:1569-1575.
Event
Preprints for Gentle Science
» MitoFit Preprints - the Open Access preprint server for mitochondrial physiology and bioenergetics
Labels:
Preprints