Difference between revisions of "SUIT protocol names"
| Line 2: | Line 2: | ||
|abbr=SUITp-Names | |abbr=SUITp-Names | ||
|description=[[File:SUIT-nomenclature.jpg|300px|right|SUIT protocols]] | |description=[[File:SUIT-nomenclature.jpg|300px|right|SUIT protocols]] | ||
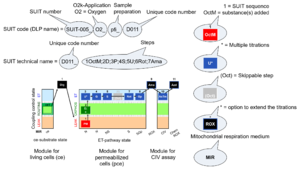
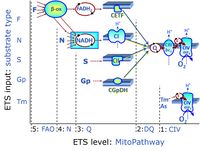
The ''' | The '''short name''' of a SUIT protocol starts with (i) the [[Categories of SUIT protocols |SUIT category]] which shows the [[ETS substrate types]] applied (controlling ETS pathway types; e.g. S, NS, FNS, FNSGp), independent of the actual sequence of titrations. (ii) A further distinction is provided in the SUIT name by listing in parentheses the [[N-pathway control |N-type substrates]] applied, again independent of the sequence of titrations, e.g. NS(GM), NS(PM), FNSGp(PGM). (iii) A sequentially selected number is added, e.g. SUIT_FNS(PM)01 (see [[Coupling-pathway control diagram]]). | ||
The ''' | The '''systematic name''' of a SUIT protocol starts with the [[Categories of SUIT protocols |SUIT category]], followed by an underline dash and the sequence of titration steps (mark names, #''X'', separated by a comma). The [[Marks in DatLab |Marks]] define the section of a [[respiratory state]] in the SUIT protocol. The [[Mark names in DatLab |Mark name]] contains the sequential number and the [[metabolic control variable]], ''X''. The metabolic control variable is the name of the preceding SUIT [[event]]. The [[MitoPedia: SUIT |MitoPedia list of SUIT protocols]] can be sorted by the short name or the systematic name (hence by SUIT protocol category. The '''[[SUIT protocol pattern]]''' is best illustrated by a [[coupling-pathway control diagram]]. | ||
|info=[[MiPNet21.06 SUIT RP]] | |info=[[MiPNet21.06 SUIT RP]] | ||
}} | }} | ||
| Line 10: | Line 10: | ||
|mitopedia concept=MiP concept, SUIT concept | |mitopedia concept=MiP concept, SUIT concept | ||
}} | }} | ||
Contributed by [[Gnaiger Erich]] 2016-02-01; edited 2016-09-05. | Contributed by [[Gnaiger Erich]] 2016-02-01; edited 2016-09-05, 2016-09-16. | ||
[[File:SUIT-catg FNSGpCIV.jpg|left|200px]] | [[File:SUIT-catg FNSGpCIV.jpg|left|200px]] | ||
| Line 18: | Line 18: | ||
::::» [[Categories of SUIT protocols]] | ::::» [[Categories of SUIT protocols]] | ||
Revision as of 12:09, 16 September 2016
Description
The short name of a SUIT protocol starts with (i) the SUIT category which shows the ETS substrate types applied (controlling ETS pathway types; e.g. S, NS, FNS, FNSGp), independent of the actual sequence of titrations. (ii) A further distinction is provided in the SUIT name by listing in parentheses the N-type substrates applied, again independent of the sequence of titrations, e.g. NS(GM), NS(PM), FNSGp(PGM). (iii) A sequentially selected number is added, e.g. SUIT_FNS(PM)01 (see Coupling-pathway control diagram).
The systematic name of a SUIT protocol starts with the SUIT category, followed by an underline dash and the sequence of titration steps (mark names, #X, separated by a comma). The Marks define the section of a respiratory state in the SUIT protocol. The Mark name contains the sequential number and the metabolic control variable, X. The metabolic control variable is the name of the preceding SUIT event. The MitoPedia list of SUIT protocols can be sorted by the short name or the systematic name (hence by SUIT protocol category. The SUIT protocol pattern is best illustrated by a coupling-pathway control diagram.
Abbreviation: SUITp-Names
Reference: MiPNet21.06 SUIT RP
MitoPedia concepts:
MiP concept,
SUIT concept
Contributed by Gnaiger Erich 2016-02-01; edited 2016-09-05, 2016-09-16.