Description
Abbreviation: ATPase (PM)
Reference: A: Determination of ATPase activity in mitochondrial preparations.
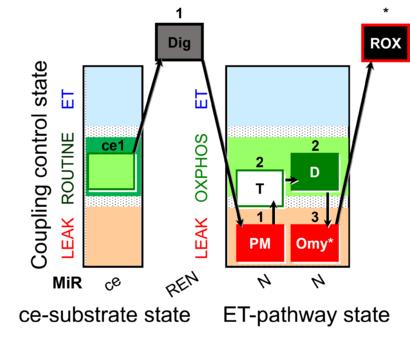
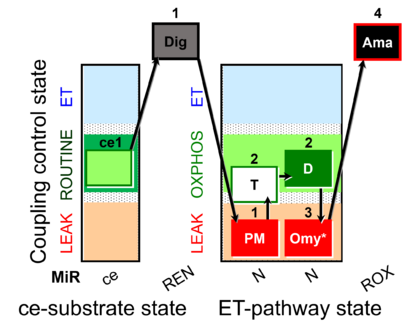
- SUIT protocol pattern: ce1;1Dig;1PM;2T;2D;3Omy;4Ama
SUIT-024 is designed to quantify the ATPase activity of mitochondrial preparations by addition of ATP (step 2T). ATPase activity in isolated mitochondria suggests contamination of the preparation by non-mitochondrial membranes. Endogenous adenylates are phosphorylated to ATP in the LEAK state, which is recycled to ADP in isolated mitochondria contaminated by ATPases, which leads to an overestimation of LEAK respiration, when comparing L(n) and L(Omy). Tissue homogenates, permeabilized muscle fibers and permeabilized cells contain all cellular membranes with high ATPase activity. The present SUIT-024 DLP file shows an application with ce-pce in the pathway category N(PM). For using this protocol with other mitochondrial preparations or substrate/inhibitor combinations, a personalized DLPU can be created.
Communicated by Garcia-Souza LF, Cardoso LHD, Gnaiger E (last update 2020-04-17)
Specific SUIT protocols
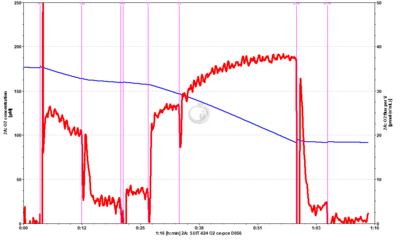
SUIT-024 O2 ce-pce D056
- SUIT-024 O2 ce-pce D056 for intact cells (ce)
Steps and respiratory states
| Step | State | Pathway | Q-junction | Comment - Events (E) and Marks (M) |
|---|---|---|---|---|
| ce1 | ROUTINE | ce1
| ||
| 1Dig | REN | ce1;1Dig
|
| Step | State | Pathway | Q-junction | Comment - Events (E) and Marks (M) |
|---|---|---|---|---|
| 1PM | PML(n) | N | CI | 1PM
|
| 2T | PML or PMP | N | CI | 1PM;2T
|
| 2D | PMP | N | CI | 1PM;2T;2D
|
| 3Omy | PML(Omy) | 1PM;2T;2D;3Omy
| ||
| 4Ama | ROX | 1PM;2T;2D;3Omy;4Ama
|
- Bioblast links: SUIT protocols - >>>>>>> - Click on [Expand] or [Collapse] - >>>>>>>
- Coupling control
- Pathway control
- Main fuel substrates
- » Glutamate, G
- » Glycerophosphate, Gp
- » Malate, M
- » Octanoylcarnitine, Oct
- » Pyruvate, P
- » Succinate, S
- Main fuel substrates
- Glossary
Strengths and limitations
- This protocol allows to determine ATPase acitivity in mitochondrial preparations. It is particularly useful as a quality control of the purity of isolated mitochondria.
- + Different combinations of substrates and inhibitors can be used depending on the specific aims, e.g.: PGM, PM, GM, MnaPM or other N-pathway substrate combinations; S or RotS for the S-pathway control state.
- + LEAK respiration is evaluated by inhibition of ATP synthase by Oligomycin.
- + Reasonable duration of the experiment.
- - If ATPase activity is actually higher than OXPHOS capacity, then the maximum ATPase activity cannot be quantified.
- - This protocol does not include Electron transfer pathway control steps.
- - CIV activity is not measured, to save experimental time.
- - Careful washing of the chamber is required after the experiment to avoid carry-over of oligomycin.
Compare SUIT protocols
References
MitoPedia concepts: MiP concept, SUIT protocol, Recommended
MitoPedia methods:
Respirometry